Standard Bayesian Optimization with FCVOpt
While FCVOpt is designed for hyperparameter optimization via fractional cross-validation, the underlying BayesOpt class can be used for standard Bayesian optimization of any black-box function—no cross-validation required.
This notebook demonstrates BayesOpt on the classic Branin function, a standard benchmark for global optimization. Unlike the CV setting, the observed objective values here are direct evaluations of the true loss function (not fold-level proxies), so we can meaningfully plot the best observed value so far and track how the optimizer converges to the global minimum.
The Branin Function
The 2-dimensional Branin function is defined as:
with standard constants \(a=1\), \(b=5.1/(4\pi^2)\), \(c=5/\pi\), \(r=6\), \(s=10\), \(t=1/(8\pi)\), over the domain \(x_1 \in [-5, 10]\), \(x_2 \in [0, 15]\).
It has three global minima, all attaining \(f^* \approx 0.3979\):
\((-\pi,\; 12.275)\)
\((\pi,\; 2.275)\)
\((9.425,\; 2.475)\)
[1]:
import numpy as np
import matplotlib.pyplot as plt
from fcvopt.optimizers import BayesOpt
from fcvopt.configspace import ConfigurationSpace
from ConfigSpace import Float
Define the Objective Function
BayesOpt expects an objective that takes a dict of hyperparameter values and returns a scalar. We wrap the Branin formula accordingly.
[2]:
def branin(params: dict) -> float:
"""Branin function. Global minimum f* ≈ 0.3979."""
x1, x2 = params['x1'], params['x2']
a, b, c = 1.0, 5.1 / (4 * np.pi**2), 5.0 / np.pi
r, s, t = 6.0, 10.0, 1.0 / (8 * np.pi)
return float(a * (x2 - b * x1**2 + c * x1 - r)**2 + s * (1 - t) * np.cos(x1) + s)
BRANIN_OPTIMUM = 0.397887
print(f"Branin global minimum: {BRANIN_OPTIMUM:.6f}")
print(f"Value at (pi, 2.275): {branin({'x1': np.pi, 'x2': 2.275}):.6f}")
Branin global minimum: 0.397887
Value at (pi, 2.275): 0.397887
Define the Search Space
We use FCVOpt’s ConfigurationSpace to declare the two continuous inputs with their standard Branin bounds.
[3]:
config = ConfigurationSpace(seed=42)
config.add([
Float('x1', bounds=(-5.0, 10.0)),
Float('x2', bounds=(0.0, 15.0)),
])
print(config)
Configuration space object:
Hyperparameters:
x1, Type: UniformFloat, Range: [-5.0, 10.0], Default: 2.5
x2, Type: UniformFloat, Range: [0.0, 15.0], Default: 7.5
Run Standard Bayesian Optimization
BayesOpt follows the same API as FCVOpt but fits a standard (non-hierarchical) GP directly on the observed (x, y) pairs.
Key arguments:
Argument |
Description |
|---|---|
|
Black-box function: |
|
Hyperparameter / input search space |
|
|
|
Local directory for MLflow experiment logs |
|
MLflow experiment name |
|
Random seed for reproducibility |
We run 30 total evaluations: 5 random initializations followed by 25 acquisition-guided steps.
[4]:
optimizer = BayesOpt(
obj=branin,
config=config,
acq_function='EI',
tracking_dir='./opt_runs/',
experiment='branin_standard_bo',
seed=123,
)
best_conf = optimizer.optimize(n_trials=30, n_init=5)
optimizer.end_run()
Number of candidates evaluated.....: 30
Observed obj at incumbent..........: 0.398315
Estimated obj at incumbent.........: 0.423703
Best Configuration at termination:
Configuration(values={
'x1': 9.4327962845392,
'x2': 2.4926768337029,
})
[5]:
print(f"Best configuration found:")
print(f" x1 = {dict(best_conf)['x1']:.4f}")
print(f" x2 = {dict(best_conf)['x2']:.4f}")
print(f" f(x1, x2) = {branin(dict(best_conf)):.6f}")
print(f" Known global minimum = {BRANIN_OPTIMUM:.6f}")
print(f" Optimality gap = {branin(dict(best_conf)) - BRANIN_OPTIMUM:.6f}")
Best configuration found:
x1 = 9.4328
x2 = 2.4927
f(x1, x2) = 0.398315
Known global minimum = 0.397887
Optimality gap = 0.000428
Optimization Progress
Internally, BayesOpt selects the incumbent at each step as the training point with the lowest GP posterior mean—not necessarily the lowest observed value. This means f_inc_obs (the observed value at the GP-based incumbent) can fluctuate non-monotonically as the GP model is updated.
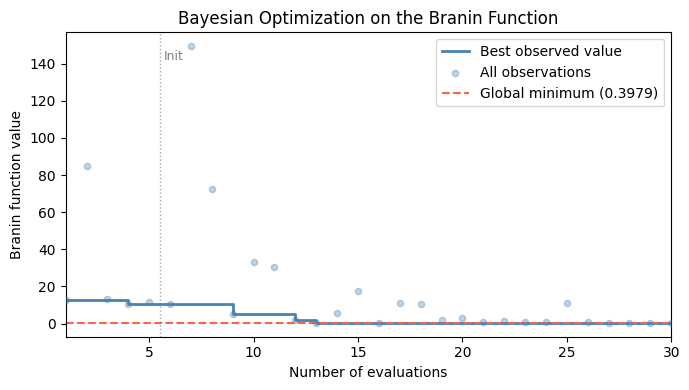
Because we are optimizing a direct, noise-free function rather than a CV proxy, a cleaner quantity to track is the best observed value so far: the running minimum over all evaluations. This is guaranteed to decrease monotonically and directly reflects how much the optimizer has improved the objective.
The plot below shows this quantity alongside all individual observations, with the known global minimum as a dashed reference line.
[6]:
observed_y = optimizer.observed_values
best_so_far = np.minimum.accumulate(observed_y)
n_evals = np.arange(1, len(observed_y) + 1)
fig, ax = plt.subplots(figsize=(7, 4))
ax.step(n_evals, best_so_far, where='post', color='steelblue', lw=2, label='Best observed value')
ax.scatter(n_evals, observed_y, color='steelblue', alpha=0.35, s=20, zorder=3, label='All observations')
ax.axhline(BRANIN_OPTIMUM, color='tomato', lw=1.5, ls='--', label=f'Global minimum ({BRANIN_OPTIMUM:.4f})')
ax.axvline(5.5, color='gray', lw=1, ls=':', alpha=0.7)
ax.text(5.7, observed_y.max() * 0.95, 'Init', color='gray', fontsize=9)
ax.set_xlabel('Number of evaluations')
ax.set_ylabel('Branin function value')
ax.set_title('Bayesian Optimization on the Branin Function')
ax.legend(loc='upper right')
ax.set_xlim(1, len(observed_y))
plt.tight_layout()
plt.show()
[ ]: